for Timeseries Forecasting
in
founder of
Stephan Sahm

- Founder of
Jolin.io
- Full stack data science consultant
- Applied stochastics, uncertainty handling
- Big Data, High Performance Computing and Real-time processing
- Making things production ready
 Jolin.io
Jolin.io
- We are based in Munich, working for Europe
- We focus on Julia consulting, high performance computing, machine learning and data processing.
- We build end-to-end solutions, including data architecture, MLOps, DevOps, GDPR, user interface, ...
- 10+ years experience in Data Science
5+ years in IT consulting
5+ years with Julia
Outline
 introduction
introduction
- Developed and incubated at MIT
- Just had 10th anniversary
- Version 1.0 in 2018
- Generic programming language
- Focus on applied mathematics
- Alternative to Python, R, Fortran, Matlab
3 Revolutions at once
Where is  used in production?
used in production?
| Industries | Pharma, Energy, Finance, Medicine & Bio-technology |
| Fields of application | Modeling/simulation, optimization/planning, Data Science, High Performance Computing, Big Data, Real-time |
Multimethods
intuitive generic programming - every function is an interface
Method specialization works for arbitrary many arguments as well as if types and functions are in different packages
Mandel-brot Example
Julia 30x faster than Python & Numpy
100% Julia versus mixture Python & C
200x200 in 10 sec
Optimising Python is generally depending on performant packages like Numpy & Numba.
better performance
=
better usage of performant packages
200x200 in 0.4 sek
Optimising Julia can be done everywhere.
better performance
=
better usage of Julia itself
Differential Equations within Neural Nets
Dynamics of population of rabbits and wolves.
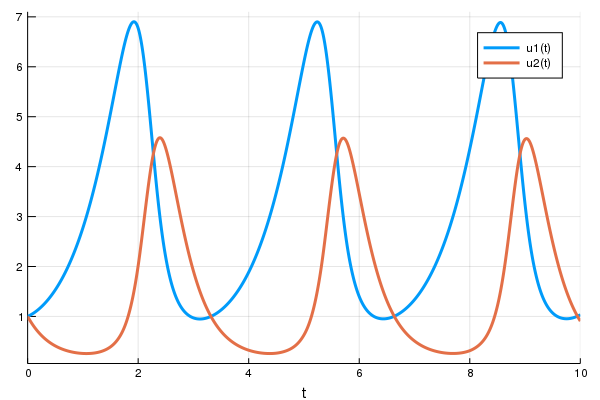
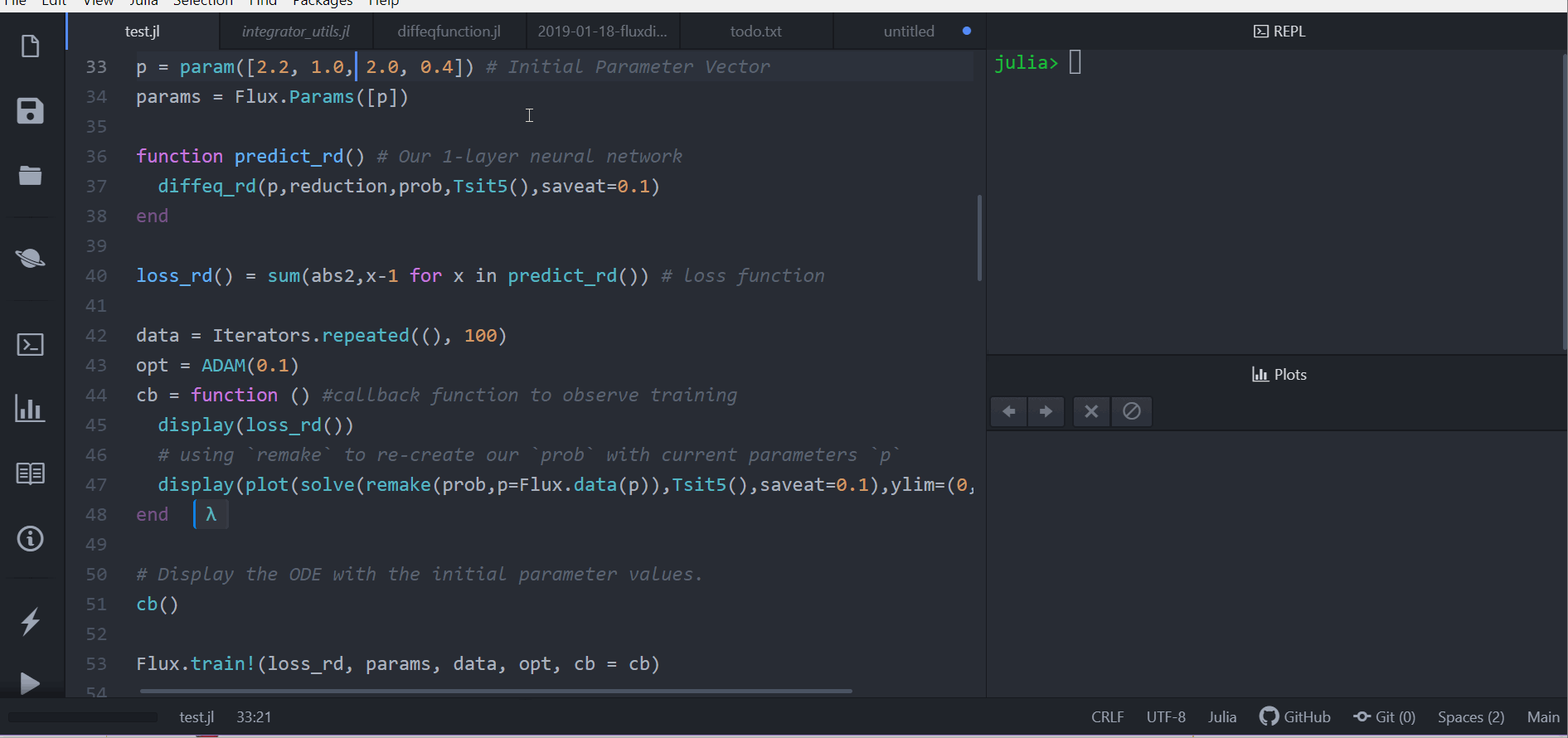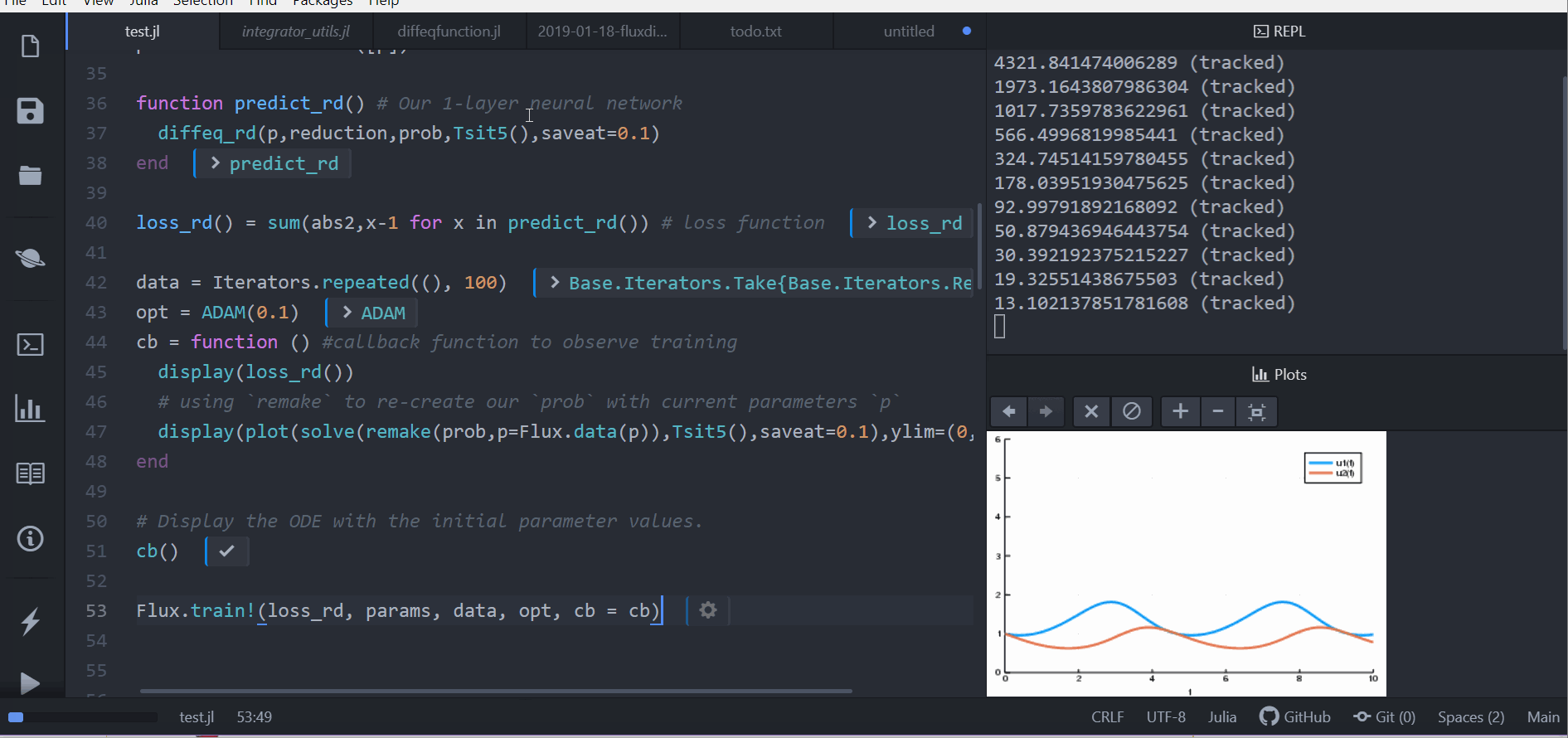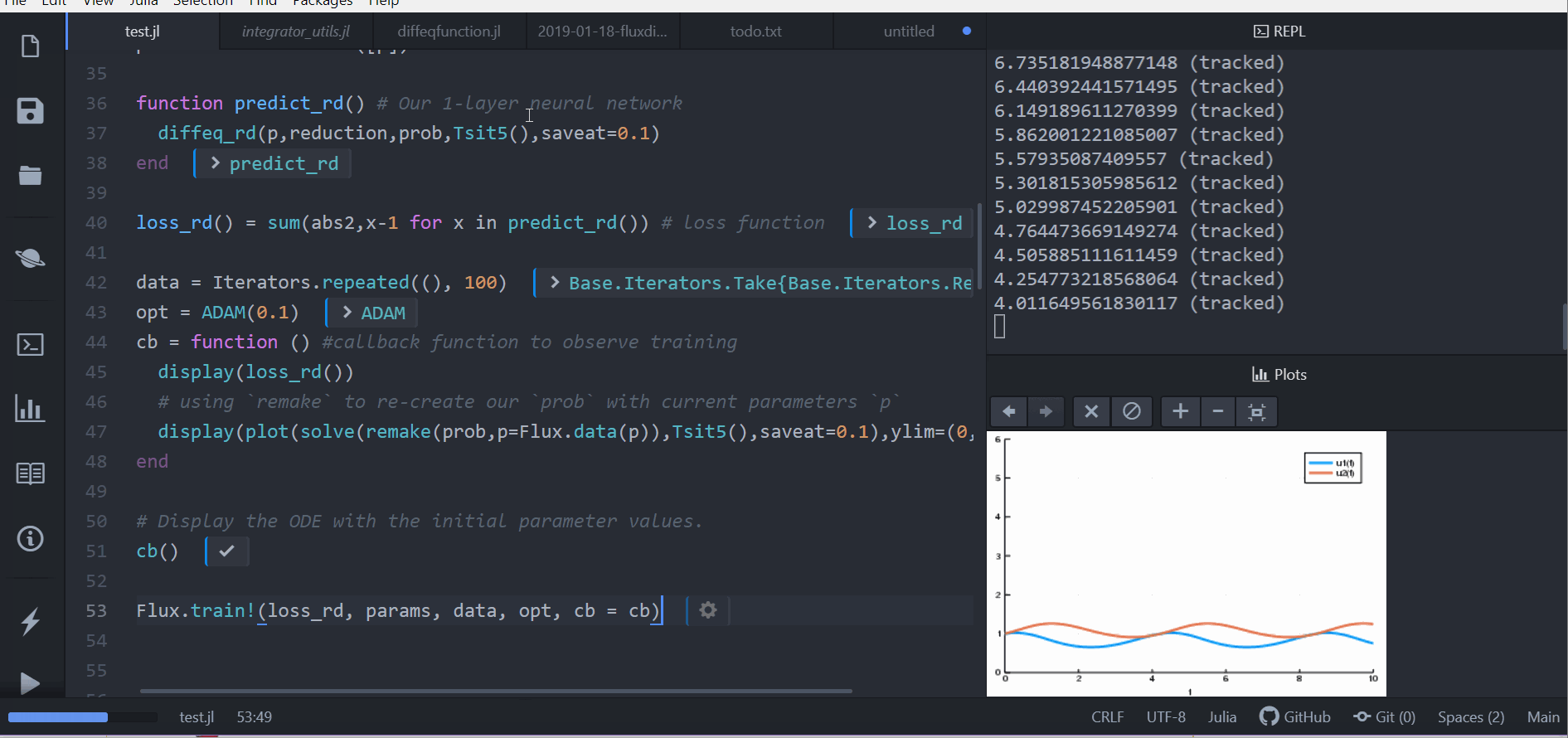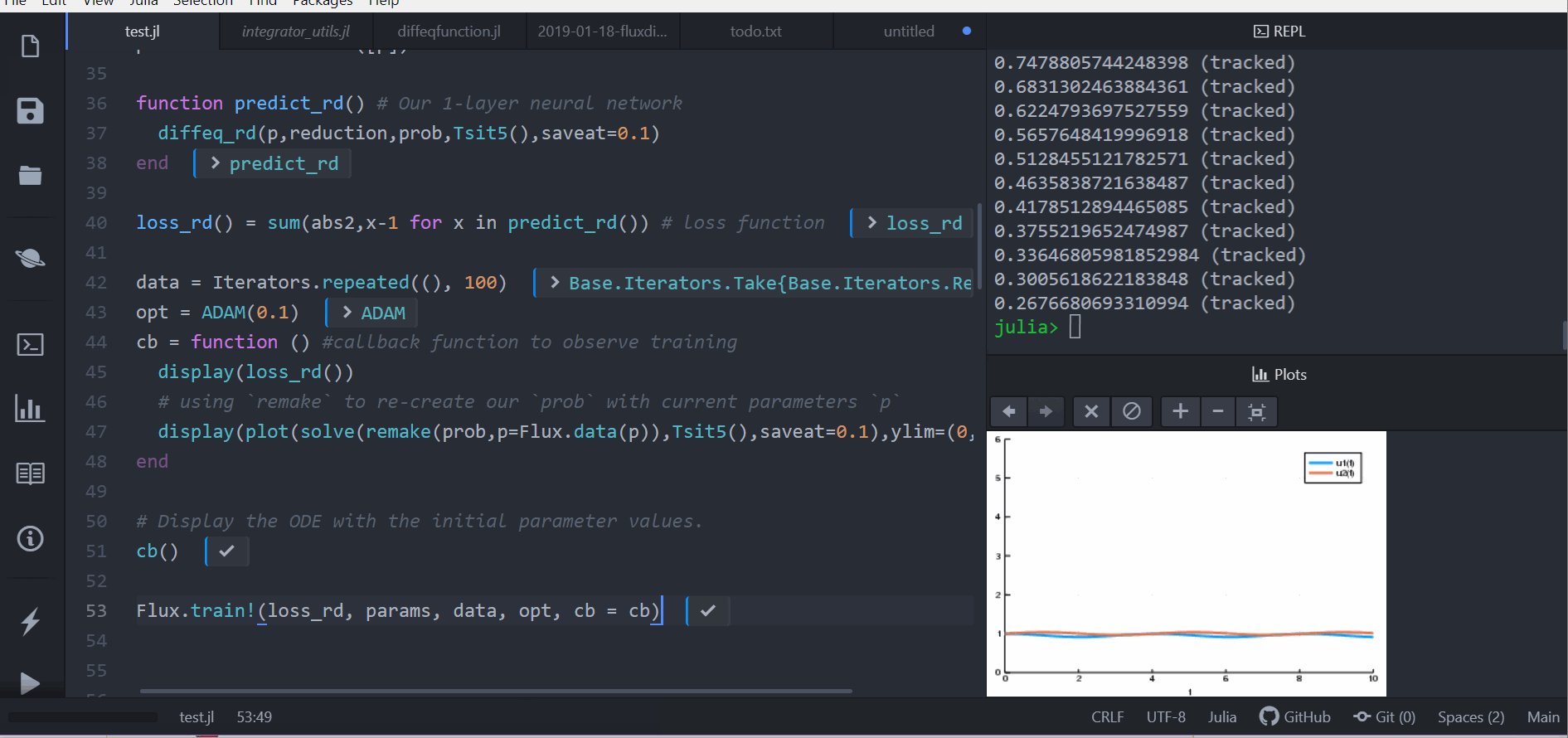
Julia model
Source: https://julialang.org/blog/2019/01/fluxdiffeq/
Putting ODE into Neural Network Framework Flux.jl
loss function (let's say we want a constant number of rabbits)
train for 100 epochs
can be part of larger network
Neural Nets within Differential Equations
Ground Truth $u^\prime = A u^3$

Model $u^\prime$ with neural network.
(multilayer perceptron with 1 hidden layer and a tanh activation function)
Source: https://julialang.org/blog/2019/01/fluxdiffeq/
plot
train

Neural Nets within Differential Equations
Alternative motivation for Neural Differential Equations: Generalization from Residual Nets.
Source: Neural Ordinary Differential Equations (Chen et al. 2019)
Residual Neural Network
discrete difference layers
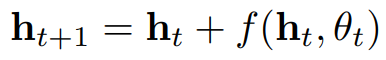
Neural Ordinary Differential Equation
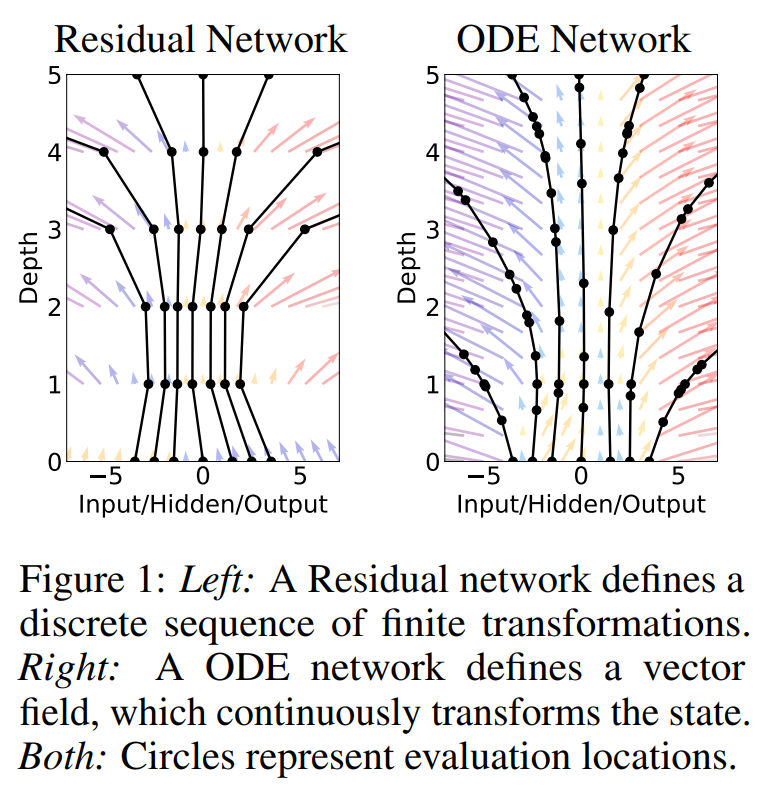
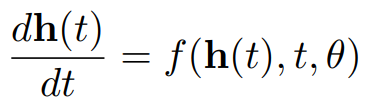

Uncertainty Learning - Bayesian Estimation and DiffEq
Let's assume noisy data

Let's assume we only have predator-data (wolves)
Probabilistic Model of the parameters
Source: https://turing.ml/dev/tutorials/10-bayesian-differential-equations/
Sample & plot

Symbolic Regression
Extract human readable formula from learned Neural Differential Equation
Benchmarks
Source: Universal Differential Equations for Scientific Machine Learning (Rackauckas et al. 2021)
Features
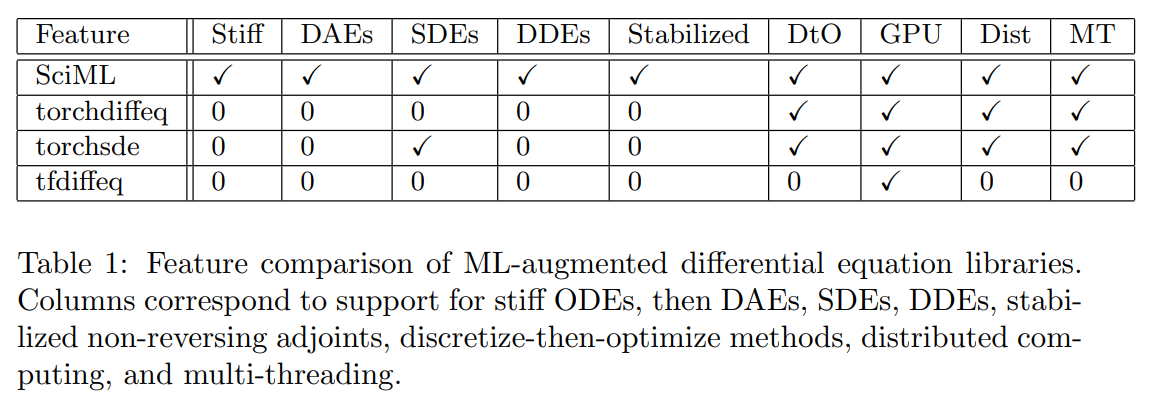
Speed
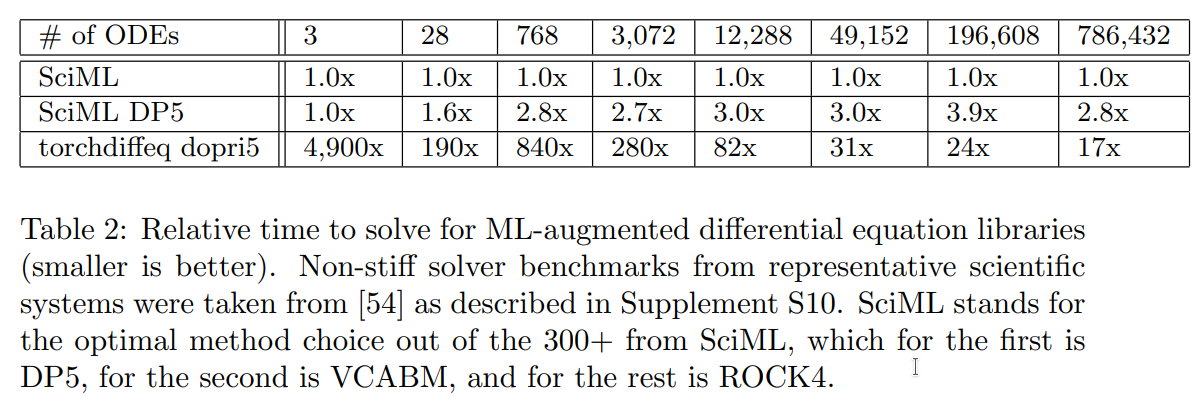
“torchdiffeq’s adjoint calculation diverges on all but the first two examples due to the aforementioned stability problems.”
tfdiffeq (TensorFlow equivalent): “speed is almost the same as the PyTorch (torchdiffeq) code base (±2%)”
Summary
- 30x-300x faster than Python
- 100% Julia
- Multimethods
- Think of an ODE as another layer for your Neural Network.
- Works with ODE, SDE, DDE, DAE, PDE.
- Model the derivative with a Neural Network.
- Generalization of Residual Networks.
- Works with ODE, SDE, DDE, DAE, PDE.
- Model randomness.
- Capture training uncertainty.
- Works with ODE, SDE, DDE, DAE, PDE.
- Extract human readable formulas.
- Widest range of features.
- Highest speed, especially for problems with viewer parameters.
Thank you for your interest
I am happy to answer all your questions. Please reach out to me or Jolin.io.
 MI4People
MI4People

